
Jill Banfield, a scientist at the University of California, Berkeley, has been poking around in disused mines for more than 10 years, studying the microbes that inhabit these dark environs. Now, together with co-researcher Brett Baker, Banfield has discovered three new nano-sized microbes living amidst the more commonplace bacteria of the mine’s interior. All three were so small – the size of large viruses – as to be virtually invisible under a microscope, and belonged to a totally new phylum of Archaea.
Banfield’s find was made possible thanks to “shotgun sequencing”, a technique which was first used to sequence the human genome in record time. “It was amazing,” said Banfield. “These were totally new and very small organisms we didn’t know how to culture with standard techniques. This shows the great promise of shotgun sequencing to profile a community of organisms without making any assumptions about what is there.” The discovery makes clear that shotgun sequencing can pick out rare organisms too small to see easily, and too novel to be plucked out by other genetic techniques.
“We were essentially looking for new stuff, and we found it in all the samples studied, though at low abundance,” explained Baker. “Shotgun sequencing is a better way to identify organisms than using other methods, like culturing or PCR [polymerase chain reaction], which can miss quite a lot of organisms.”
The organisms in the mine sludge obtain their energy by oxidizing iron (generating rust) and in the process create sulfuric acid and dissolve pyrite to release more iron and sulfur. This self-sustaining process creates the acidic drainage that pollutes creeks and rivers, including those around the researchers’ study site, the Richmond Mine at Iron Mountain, California.
Shotgun sequencing involves homogenizing the organisms in a sample of the sludge, isolating the combined DNA and breaking it into lots of random strands. Each strand is then sequenced, and a powerful computer is used to find overlaps so that the pieces can be properly reordered. This process identified five separate genomes that corresponded to five bacteria and Archaea – four of them uncultivated at that time, though closely related to known microbes.
 Baker then probed the gene fragments more thoroughly to turn up three Archaea from a totally unknown group, probably representing a new phylum among the several dozen known phyla of Archaea. They fall within a large class of microbes known as thermophiles, which are Archaea that live in warm and even scalding conditions. Many of these thermophiles have been recovered from hydrothermal vents in the deep mid-ocean ridges, where lava boils up between continental plates.
Baker then probed the gene fragments more thoroughly to turn up three Archaea from a totally unknown group, probably representing a new phylum among the several dozen known phyla of Archaea. They fall within a large class of microbes known as thermophiles, which are Archaea that live in warm and even scalding conditions. Many of these thermophiles have been recovered from hydrothermal vents in the deep mid-ocean ridges, where lava boils up between continental plates.
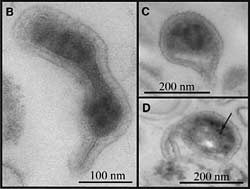
Once Baker had found gene segments (ribosomal RNA) from three Archaea, he was able to fish the microbes out of the slime soup and found that they were extremely small, around 200 nanometers in diameter, the size of large viruses. Bacteria average about five times this diameter. These could be the smallest organisms ever found, though Baker needs to culture them before confirming this. Because they’re so small, however, they may not be free-living.
“We’re not sure they can live independently, whether they have enough genes to fend for themselves, but instead are symbiotic with another organism or are feeding off another organism,” Baker said. Baker now is trying to find the right conditions for these Archaea to thrive in a culture dish. For now, he has dubbed them ARMAN-1, -2 and -3, for Archaeal Richmond Mine Acidophilic Nanoorganisms.
Related Articles
Mayo Clinic Zooms In On Nanobacteria
Nanobacteria A Problem For Space Travellers, Says NASA
More Evidence For NanoBacteria Say Researchers
Airborne Nanobacteria Spread Disease, Create Rain








Comments are closed.